Cutevariant
A standalone and free application to explore genetics variations from VCF file
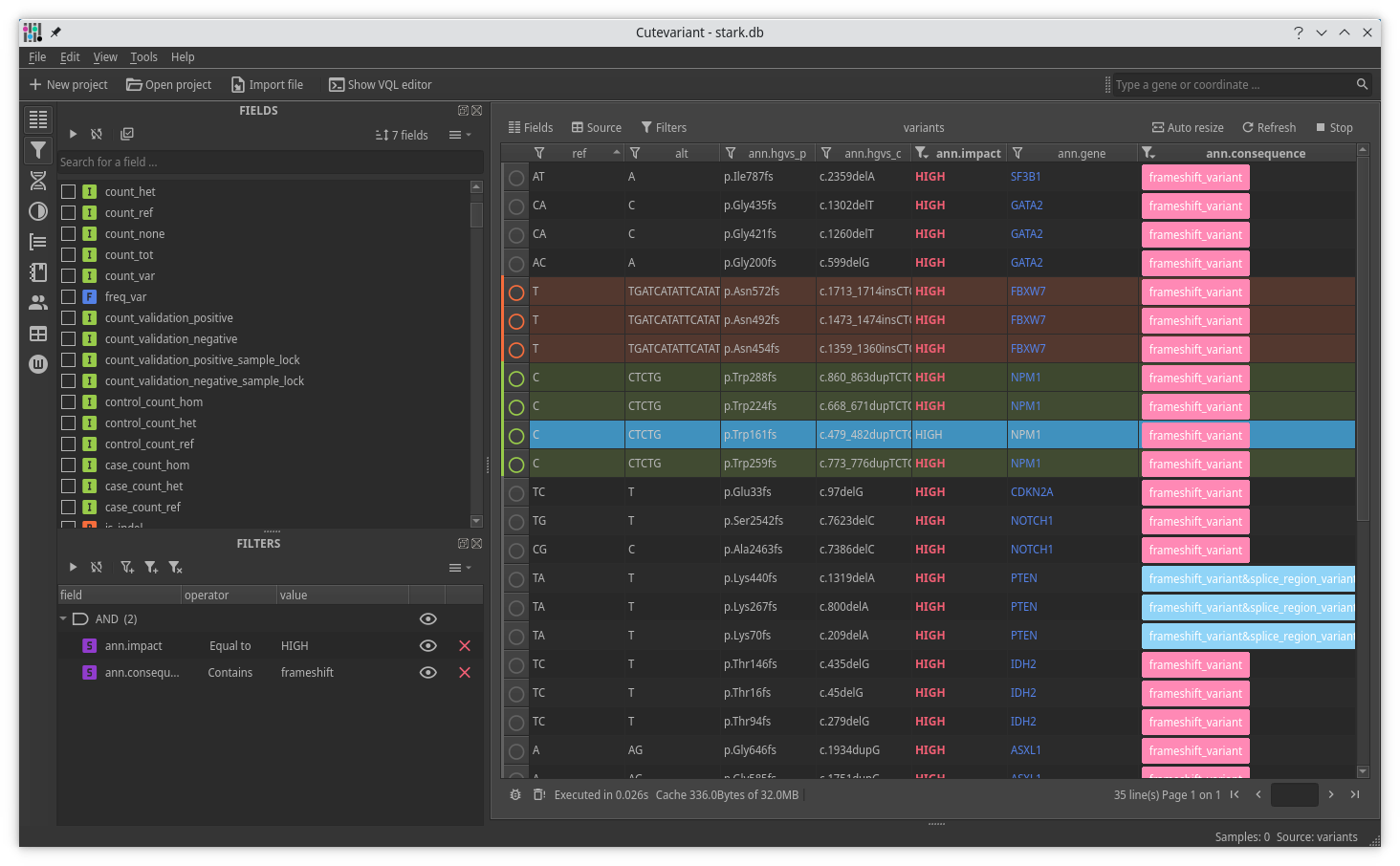
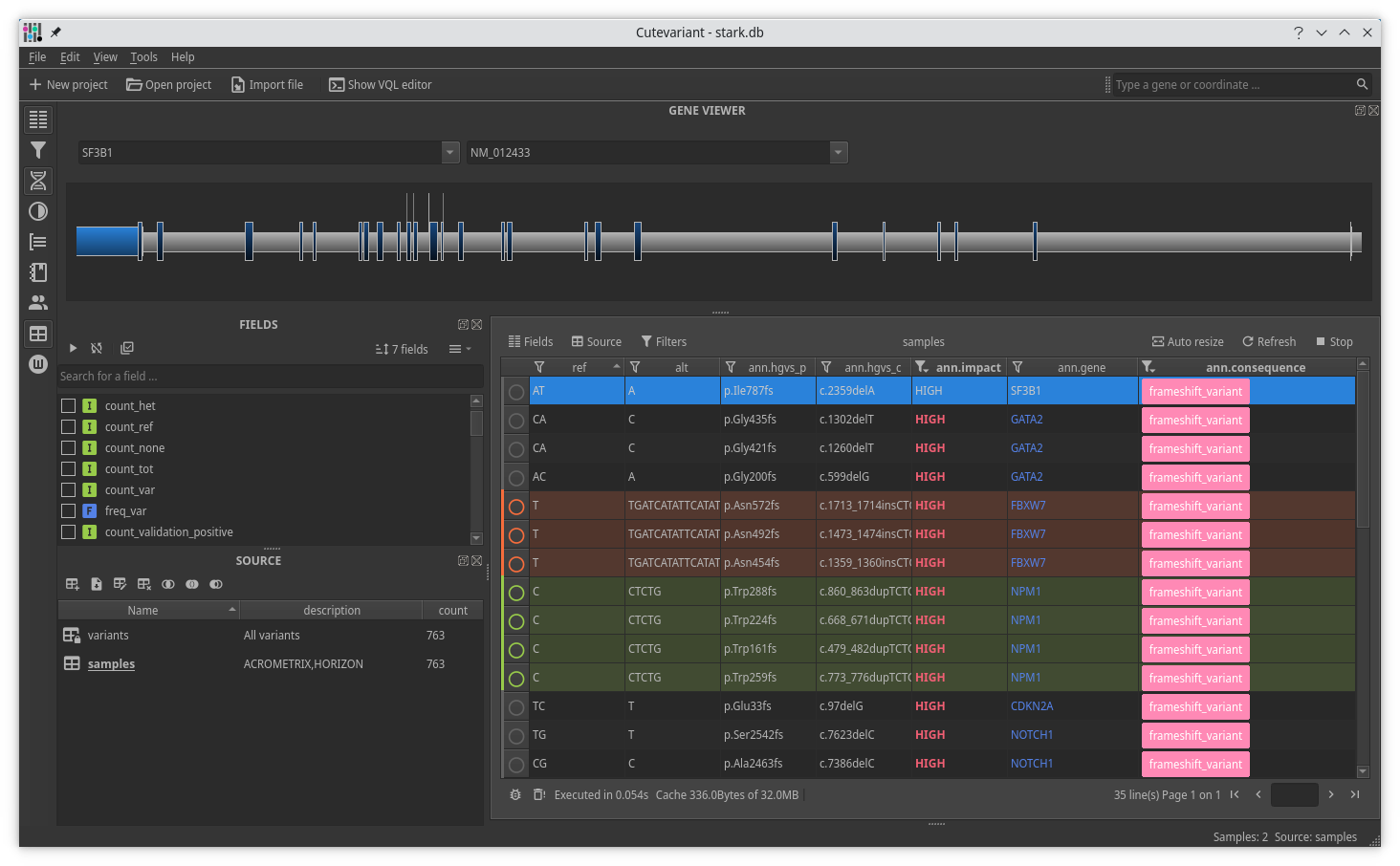
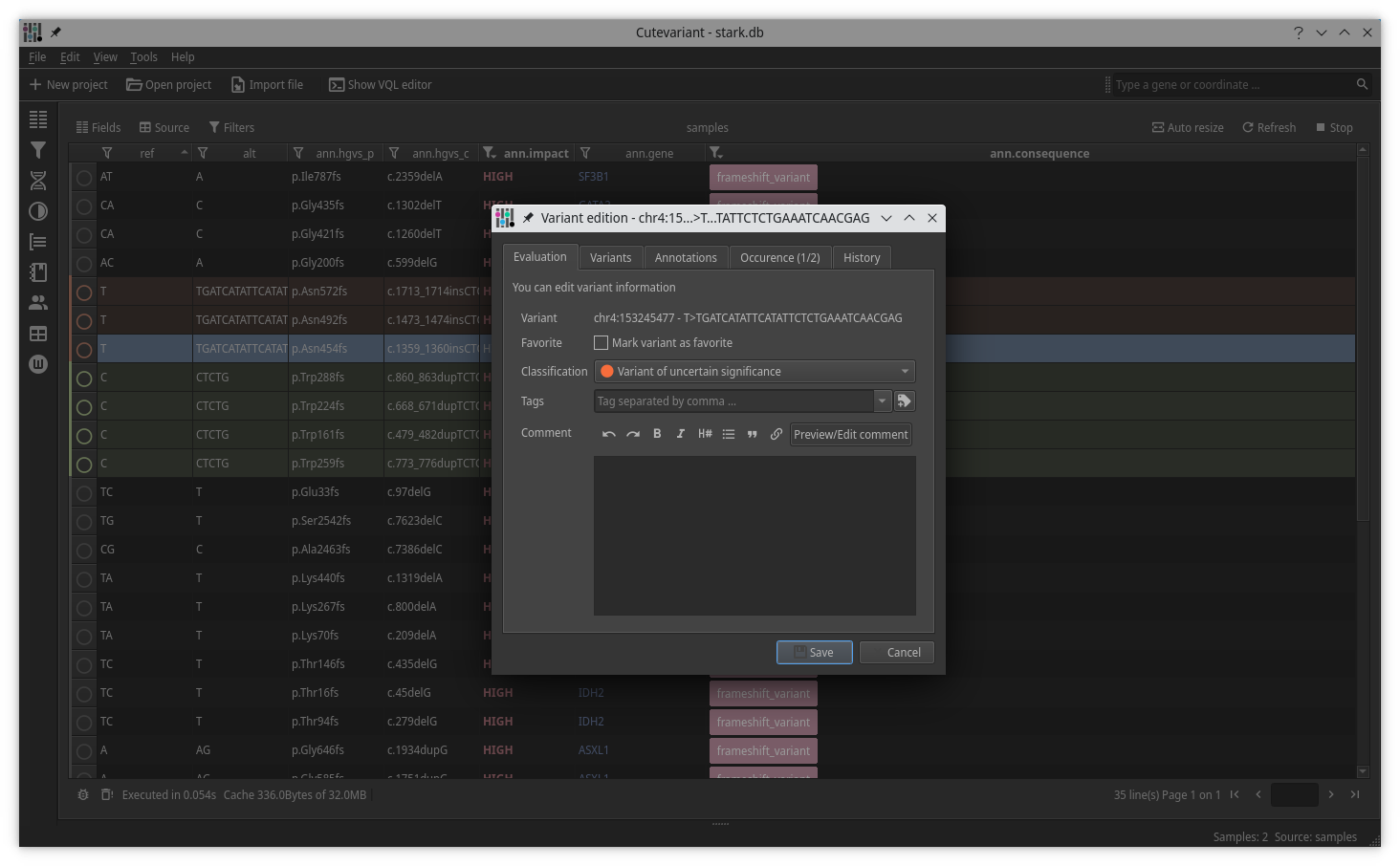
Cutevariant is a cross-plateform application dedicated to maniupulate and filter variation from annotated VCF file. When you create a project, data are imported into an sqlite database that cutevariant queries according your needs. Presently, SnpEff and VEP annotations are supported. Once your project is created, you can query variant using different gui controller or directly using the VQL language. This Domain Specific Language is specially designed for cutevariant and try to keep the same syntax than SQL for an easy use.
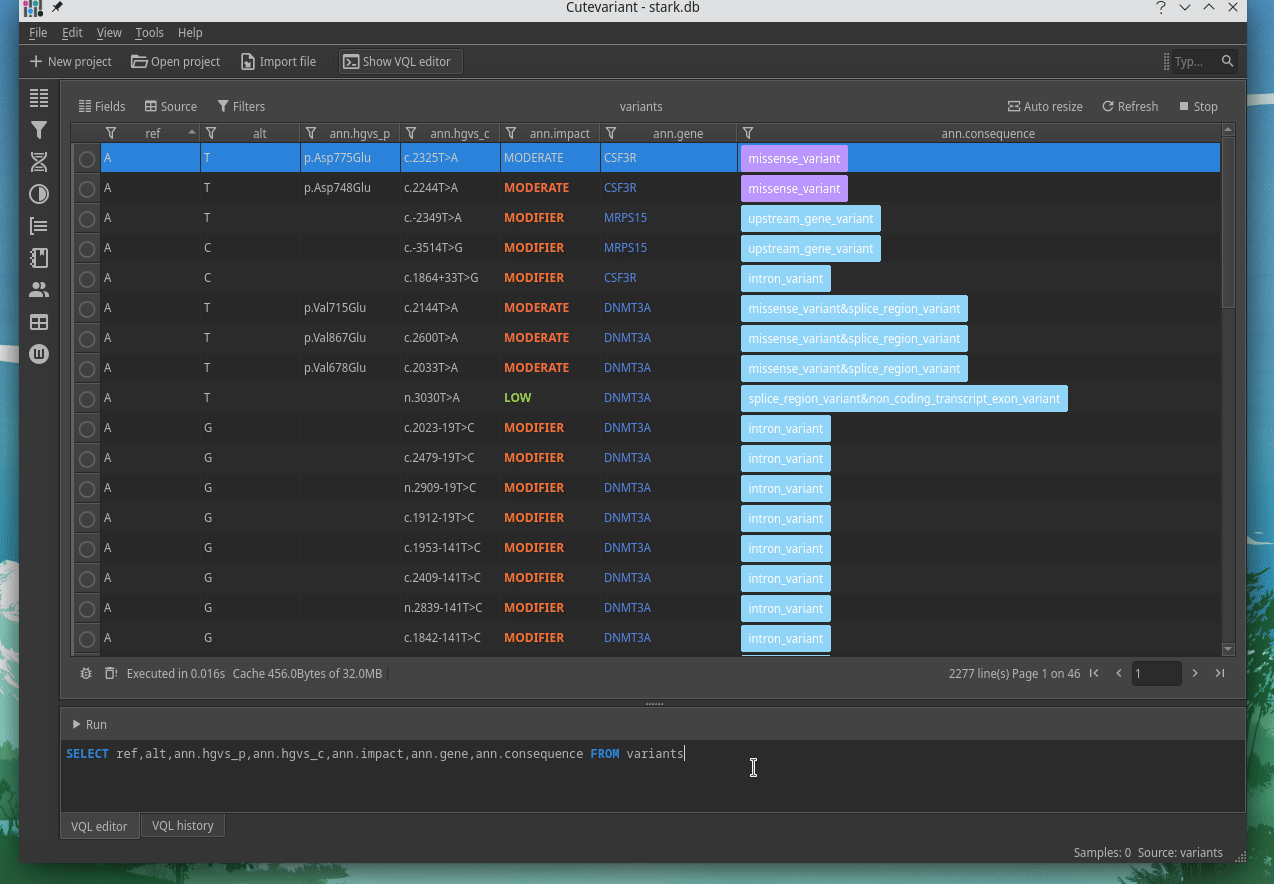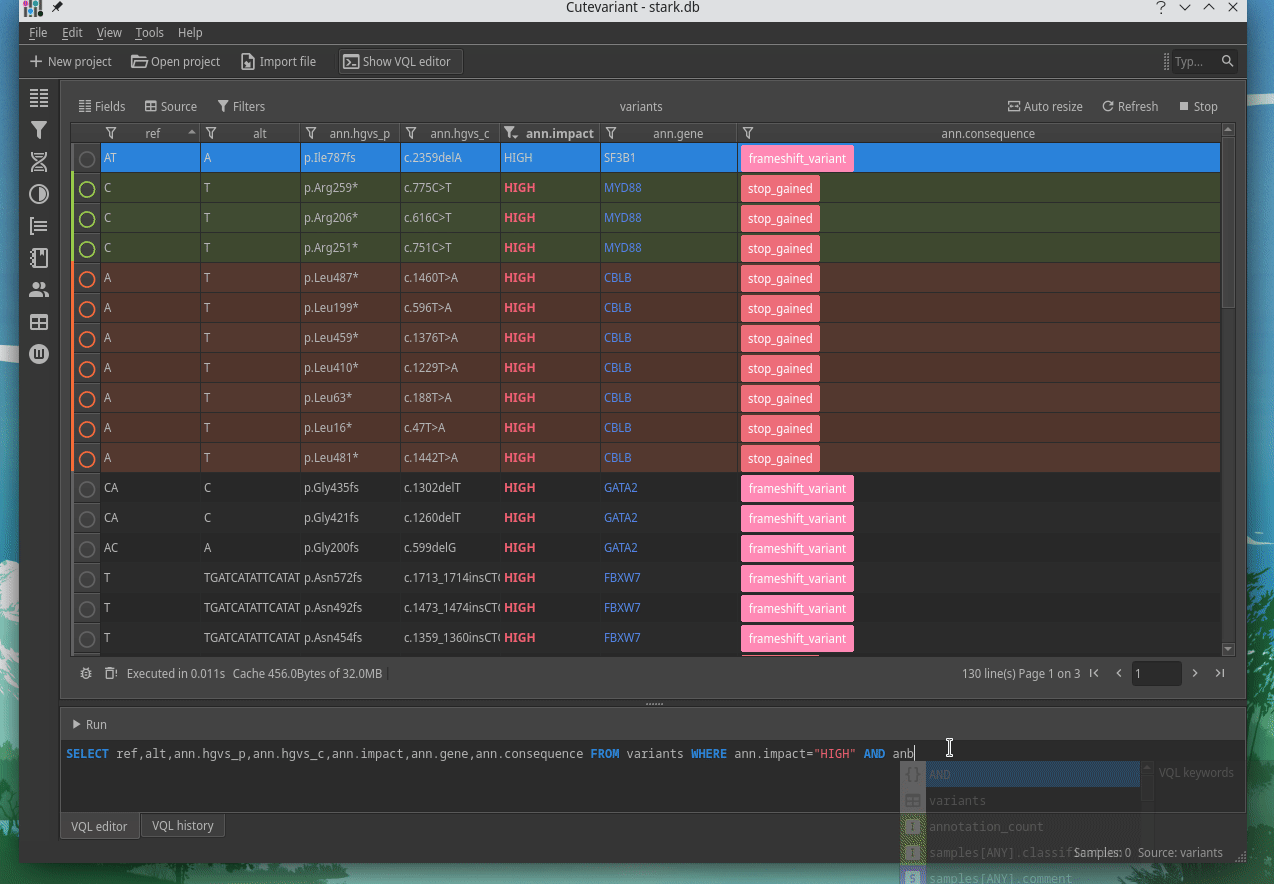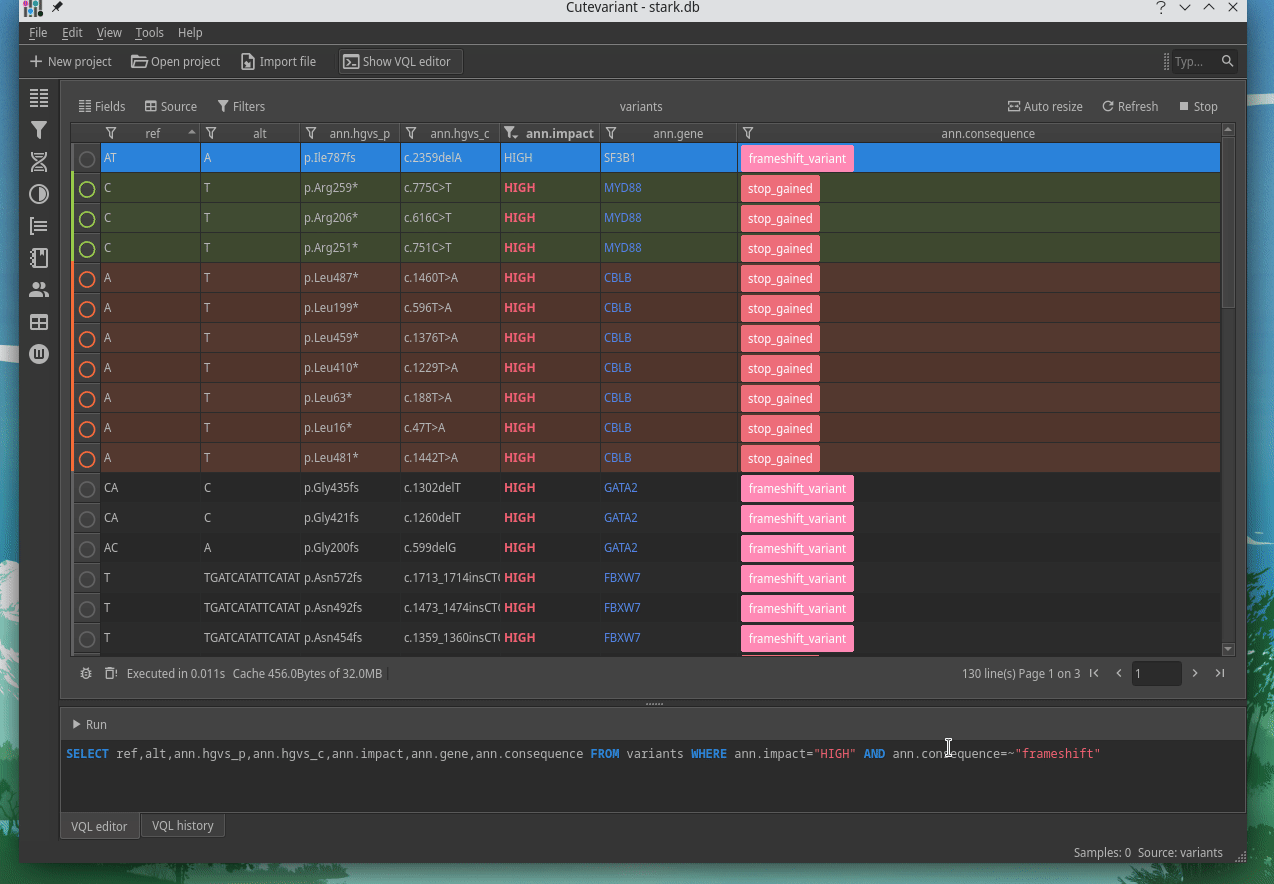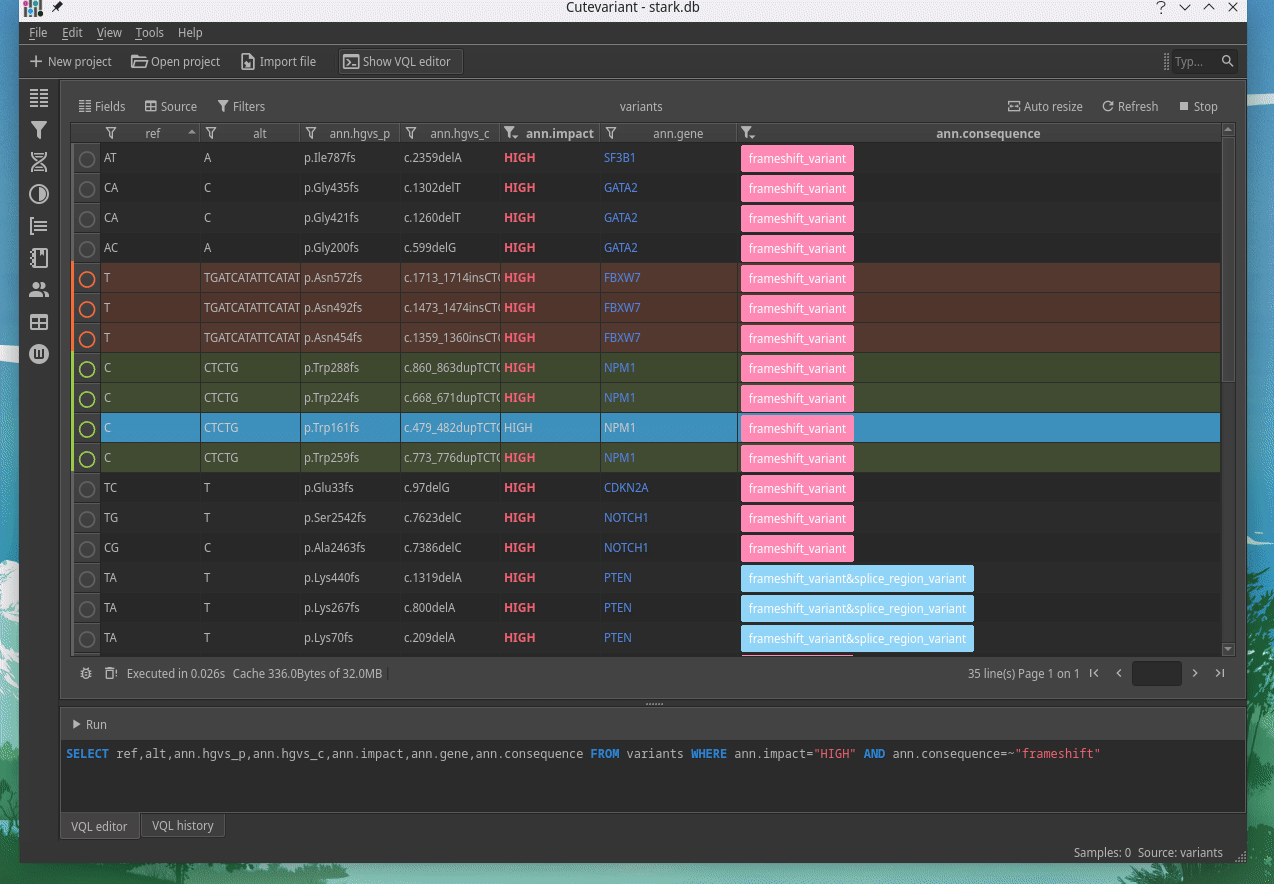
 |
 |
 |
Installation
Windows
Standalone binary are available for windows:
PyPi
Cutevariant is avaible from Pypi :
pip install cutevariant # install
python -m cutevariant # run
From source
- Python 3.6 or newer is required
git clone https://github.com/labsquare/cutevariant.git
make install_deps
make run
Usages
You can follow this tutorial to familiarize yourself with cutevarant.
Usage examples
The VQL langage specification is available here :
VQL specification
Contributions / Bugs
Cutevariant is a new project and all contributors are welcome
Issues
If you found a bug or have a feature request, you can report it from the Github isse trackers.
Create a plugin
Documentation to create a plugin is available here
Chat
You can join us on discord. We are speaking french right now, but we can switch to english.
Licenses
This program is free software: you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation, either version 3 of the License, or (at your option) any later version.
This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details.
You should have received a copy of the GNU General Public License along with this program. If not, see https://www.gnu.org/licenses/gpl-3.0.txt.